Spectral Imaging
Introduction | Analysis - Deconvolution and Classification | Information Content
Pure Component (Endmember) Estimation | Calibration and Correction
Spectral Imaging Introduction
Spectral imaging combines two disciplines - spectroscopy and photography - to sample image data at many wavelength bands. In general, spectral imaging is separated into either multispectral (< 20 wavelength bands sampled) or hyperspectral (> 20 wavelength bands). However, a more robust definition may be that multispectral imaging samples data from multiple discrete, but non-contiguous wavelengths, while hyperspectral imaging samples sufficient data to reconstruct a contiguous spectrum, over a given spectral range. Originally developed by NASA and the Department of Defense for remote-sensing applications, spectral imaging is now used for biological applications. In particular, spectral imaging approaches have been utilized for separating signals from multiple fluorescent labels in cells or tissues. Our research in biomedical spectral imaging makes use of the developments in remote sensing, and applies them to specific biomedical problems. For example, instead of using spectral information to locate a camouflaged tank, spectral absorbance or fluorescence can be used to track a 2nd-messenger signal in a single cell or to locate a pre-cancerous lesion of the airway or colon. For dermal, superficial, and endoscopically-accessible tissues, spectral imaging can be implemented in vivo.
A key advantage of hyperspectral imaging is the ability to simultaneously measure many fluorophores in a field-of-view. This stands in contrast to traditional fluorescence microscopy, where a series of bandpass filters are used to isolate signal from one-to-three fluorophores at a time. Several technologies are available to acquire hyperspectral image data:
- Tunable filters, which scan wavelength-bands sequentially
- Diffraction gratings and prisms, which disperse light into constituent wavelengths
- Interferometers, which use constructive and destructive interference
The resulting image set, sometimes referred to as a “spectral cube” is then analyzed to extract wavelength-dependent features, such as the relative abundance of each molecular species (referred to as an endmember). This powerful approach can be used to identify a single fluorescent label amongst many, even if the labels have highly-overlapping spectra.
Work in our laboratories has focused on developing new technologies for spectral imaging and on applying spectral imaging to understand cells and tissues at a molecular level. We have developed new technologies for hyperspectral imaging that have allowed 2-3 orders of magnitude increases in detection sensitivity and imaging speed (see Technology Development for more details). We have also new applications for using 5D microscopy (3D + spectral + timelapse) for monitoring cAMP and other 2nd messenger signals in 3 dimensions over time in single cells (see Second Messenger Signaling for more details). Our future work will be to further expand these capabilities, including the ability for very high-speed 5D microscopy, simultaneous measurements of multiple events in signaling cascades, intravital microscopy (especially intravital FRET and 2nd messenger signaling approaches), and clinical translation of imaging technology.
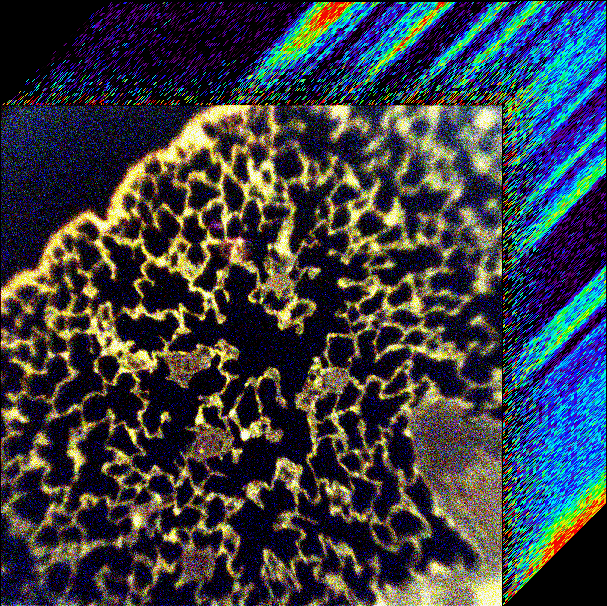
Spectral imaging utilizes spectral filtering technology to scan, or filter, wavelengths to form a contiguous spectrum. The result is a 3-D image cube, with the third dimension is each wavelength scanned. This image shows the natural fluorescence (autofluorescence) from excised rat lung tissue. The fluorescence excitation was scanned from 360-550 nm in 5 nm increments using a tunable filter apparatus connected to a widefield fluorescence microscope.
Spectral Analysis - Deconvolution and Classification
Spectral deconvolution is the process of estimating the percent contribution of individual component spectral to an overall (total) measured spectrum. In the spectral imaging field, spectral deconvolution is referred to as a sub-pixel process, as it seeks to estimate the amount (abundance) of each pure component in a given pixel. Linear unmixing is one of the most common approaches to spectral deconvolution. Linear unmixing assumes that each of the pure species (specified a priori) contributes linearly to the total spectrum.
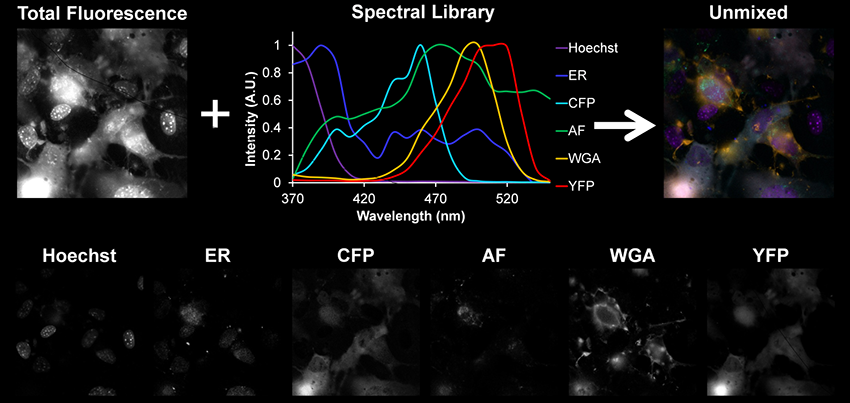
Nonnegatively-constrained linear unmixing of 6 spectral components. Pulmonary microvascular endothelial cells were labeled with Hoechst 33342 (nuclear stain), ER Tracker (endoplasmic reticulum label), a CFP-Epac-YFP FRET reporter that indicates cAMP concentration and WGA (Wheat Germ Agglutinin - membrane label). Cellular autofluorescence was also identified (labeled AF).
Spectral Information Content
One significant limitation of spectral imaging is the decrease in signal intensity compared to a non-spectral system. This is due to the dispersion of light needed to separate and detect individual wavelengths. If the spectrum is viewed as a fixed distribution (histogram), the decrease in signal intensity in a spectral imaging system is analogous to the decrease in the number of samples per bin that occurs when the number of bins increases. This dispersion of light in spectral imagers produces lower signal-to-noise ratios and/or necessitates higher image acquisition times. For live-cell imaging of dynamic events or for samples that photobleach rapidly, these increased acquisition times can be prohibitive. Thus, it is necessary to develop methods to optimize spectral imaging systems before spectral techniques can be applied to light-sensitive or high speed live cell imaging.
We are researching methods to measure the information content of spectral image sets based on the qualities of the spectral distribution (histogram) of each pixel. For example, histogram dissimilarity metrics such as Bhattacharya Distance and Earth Mover’s Distance describe the difference between two histograms (spectra) based on each spectrum as a continuous distribution. We have found these metrics to be much more sensitive at measuring subtle differences in spectral signatures than more traditional metrics, such as Spectral Angle Mapper, which view a spectrum as a series of unrelated measurements at different wavelengths. Using these metrics, we can quantitatively compare the quality of spectral information collected using different equipment and acquisition settings. By selecting parameters that maximize the information content and minimize the total acquisition time, we can optimize a spectral imaging system for a given assay. The development of optimization algorithms is a critical next step in developing multi-dimensional spectral imaging techniques that can be applied to high-speed live cell imaging, and more sophisticated assays, such as Förster resonance energy transfer and fluorophore multiplexing.
Pure Component (Endmember) Estimation
For spectral classification and unmixing, a training set is usually defined that contains all of the individual component (endmember) spectra that are expected to occur within a given biological sample. Often, a structure called a spectral library will be defined as well, as a repository of all known (predefined and user-collected) spectra. One key difference between remote sensing and biological spectral imaging is that in remote sensing applications, the end-member spectra are often well characterized, whereas in biological imaging, this may not necessarily be the case. This is due to several variables:
- The physical components that are generating contrast (cellular and extracellular proteins, macromolecules, nucleic acids, etc.) are generally smaller than the resolution limit of the imaging system. Usually many of these components are present (mixed) in any given pixel, resulting in an inability to identify a single pixel, or region of pixels, as a pure component.
- Although individual components and contrast agents (fluorophores) can be isolated and spectrophotometrically analyzed, this measured spectrum may be different from the real spectrum that these molecules present when in a biological sample. This is because many of these molecules are sensitive to environmental changes (pH, reactive oxygen species, etc.), that are usually not present in an isolated sample, and because the instrument used to measure individual component spectra is often a different instrument (such as a spectrophotometer) from the instrument used to acquire the spectral image sets.
Even when the same instrument is used to acquire both isolated component spectra and to perform the biological imaging assays, different acquisition settings are often used. Some changes (such as integration time and gain of the detector) may have little effect on classification or linear unmixing, if normalized end-members are used. However, changes in settings that modify the optical pathway often result in significant modification of the measured spectra. For example, changing the dichroic beamsplitter or emission filter between collecting isolated component spectra and performing the actual imaging assay will result in a pure component spectrum that is different from the spectrum of this component in the actual sample, and often unsuitable for use as an endmember for classification or unmixing. In the best case, some of these effects can be cancelled by unmixing the transmission curve of these different filters from the measured spectrum. Changes in the spectral dispersion element (switching between multiple gratings, or binning multiple wavelengths) will also result in endmember spectra that are different from the corresponding component spectra in the sample. Up- or down-sampling may be used to try to match endmember and sample spectral data. However, if up-sampling is performed (because fewer wavelength points were collected for the end-member spectra than for the sample spectra), sharp spectral characteristics may still be left unresolved. If a laser-based excitation source is used with a grating-based spectral detector, a physical notch is often placed in front the detector, at a location corresponding to the wavelength of the laser-line. If the location of this notch is changed between collecting isolated component spectra and the sample image set (for example, if only one laser line is used to acquire each endmember spectra, but several laser lines are used in imaging the unknown sample), this will also result in a mismatch between a component's spectrum in the training set and the spectrum of the same component in the sample.
Spectral Calibration and Correction
Wavelength calibration and flat-field correction are necessary steps in most spectral imaging systems. For systems with reproducible wavelength response, wavelength calibration typically only needs to be performed periodically.
Wavelength Calibration
While many emission standards can be used for wavelength calibration, Lerner and Zucker (Cytometry Part A, 62A:8-34, 2004) present a standard calibration method using a multi-ion discharge lamp (MIDL). The MIDL contains a mixture of gases and presents emission peaks that reproducibly occur at given wavelengths. The MIDL emission spectrum is first measured using a spectrometer and then measured using the spectral imaging system. The peak locations are identified from both systems, and the spectral imaging system is corrected to match the spectrometer measurement.
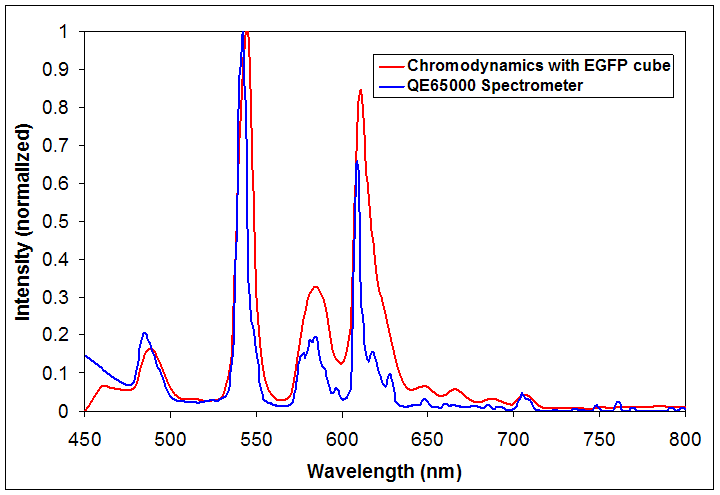
The emission spectrum of a multi-ion discharge lamp as measured using a high-sensitivity spectrometer (blue) and a spectral fluorescence microscope (red). While the spectral microscope shows greatly reduced spectral resolution, the wavelength calibration is correct to within the step size (5 nm).
Spectral Intensity Correction
Spectral intensity correction accounts for the transfer function of the spectral imaging system. When performed with appropriate controls, it can also be used to remove instrument offset, stray light, and filter bleed-through. Depending on the significance of these contributions, intensity correction can provide a significant improvement in the signal quality of the spectral image. Intensity correction is necessary when performing spectral analysis using end-members acquired under different experimental conditions or stored in a generic/manufacturer-supplied library.
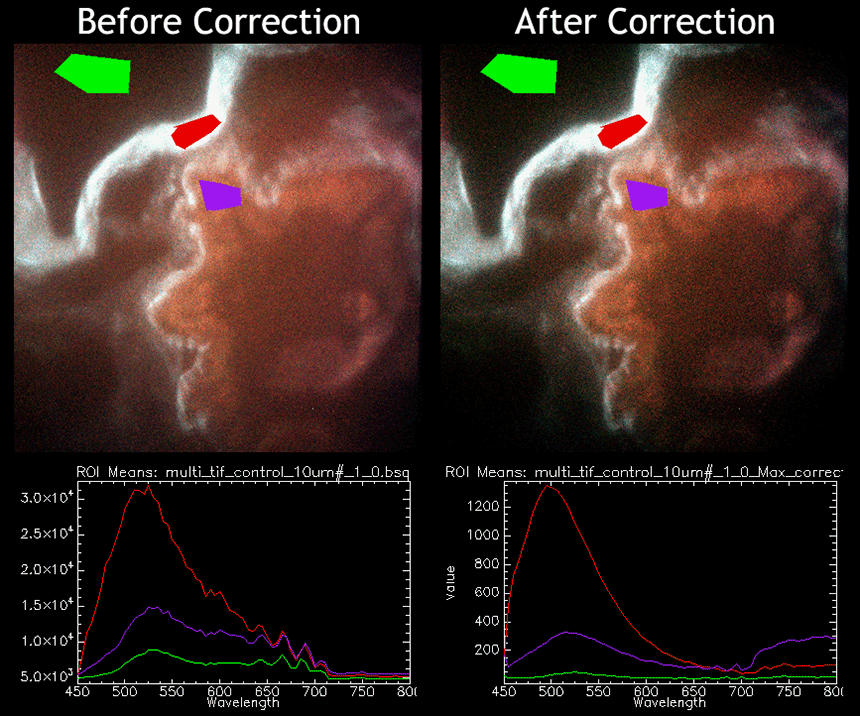
Spectral image of a 10 um unlabeled lung cryoslice showing significant improvement in spectral quality with flat-field correction. Bleed-through of the excitation source (Xe arc lamp) is clear at 530 nm and at 600 nm, and has been removed in the corrected image set. Oscillations between 600-700 nm are due to the layering of the dichroic beamsplitter and have been compensated for in the corrected image set. General background offset has been reduced. (Image taken using a custom inverted-microscope setup and acousto-optic tunable filter system (Chromodynamics); 40X-oil objective)
